Iwanaga Lab/Research Center for Infectious Disease Control Department of Parasitology
Malaria is a mosquito-borne infectious disease and is caused by infection with Plasmodium species. Its burden exceeds 200 million infections every year, resulting in more than 400,000 deaths annually. Plasmodium parasites have the complex life cycle between host animals and mosquitos. They specifically express the genes at each developmental stage during the life cycle and those stage-specific genes are essential for invasion, parasitizing, and multiplication. Our research interest is how the parasite regulate the gene expression stage-specifically. To elucidate it, we focus on the sequence-specific transcriptional factor and characterize them by the genetic engineering technique and the next generation sequencing . The expected result will be useful for understanding the molecular basis of the parasite' s life cycle, but also for exploring the drug target and vaccine antigens.
- Transcriptional regulation of Plasmodium parasites
Plasmodium parasites have the complex life cycle between host animals and mosquito vector (see: https://www.cdc.gov/ malaria/about/biology/index.html). In the course of the life cycle, the parasite change their morphology dramatically and infect specifically various cells of host animal and vectors. The
stage-specific gene expression allow the morphological change and acquirement of infectivity to cells and is thus essential for the completion of this complex life cycle. The transcriptional regulation plays an important role in this stage-specific gene expression. However, the transcriptional factors had not been identified even after completion of whole genome sequencing, the mechanism of transcriptional regulation was not elucidated. We firstly demonstrated that the Apetala2 (AP2) protein family is the sequence-specific transcriptional factor of Plasmodium parasites. AP2 transcription factors express stage specifically, and further directly and comprehensively controls the transcription of all genes involved in stage formation at each developmental stage. Our group now attempt to identify target genes of all AP2 transcription factors using next generation sequencing technology, such as chromatin-immunoprecipitation sequencing. Based on the obtained information, we attempt to elucidate mechanism of transcriptional regulation for entire life cycle.
- From Plasmodium artificial chromosome to synthetic Plasmodium parasites
In the previous study, we generated Plasmodium artificial chromosome (9.0 kbp), which consists of centromere, telomere and replication origin. It segregates into daughter parasites with more than 99.9 % efficiency during nuclear division and is maintained as single copy by the function of cent
romere. In addition, its telomere functions properly, which protects the ends. We further combined artificial chromosomes of Plasmodium parasite and budding yeasts and generated a shuttle artificial chromosome, which can behave like actual chromosomes in both living organism. We now attempt to synthesize parasite' s genome in budding yeast and to transplant the resultant synthetic genome into Plasmodium parasite. The transgenic parasites. in which synthetic genome will be transplanted. will be first synthetic eukaryotic cells and will be utilized for synthetic biological research. Furthermore, it will be utilized for designing artificial attenuated parasites, which will be safety live vaccine for malaria.
-
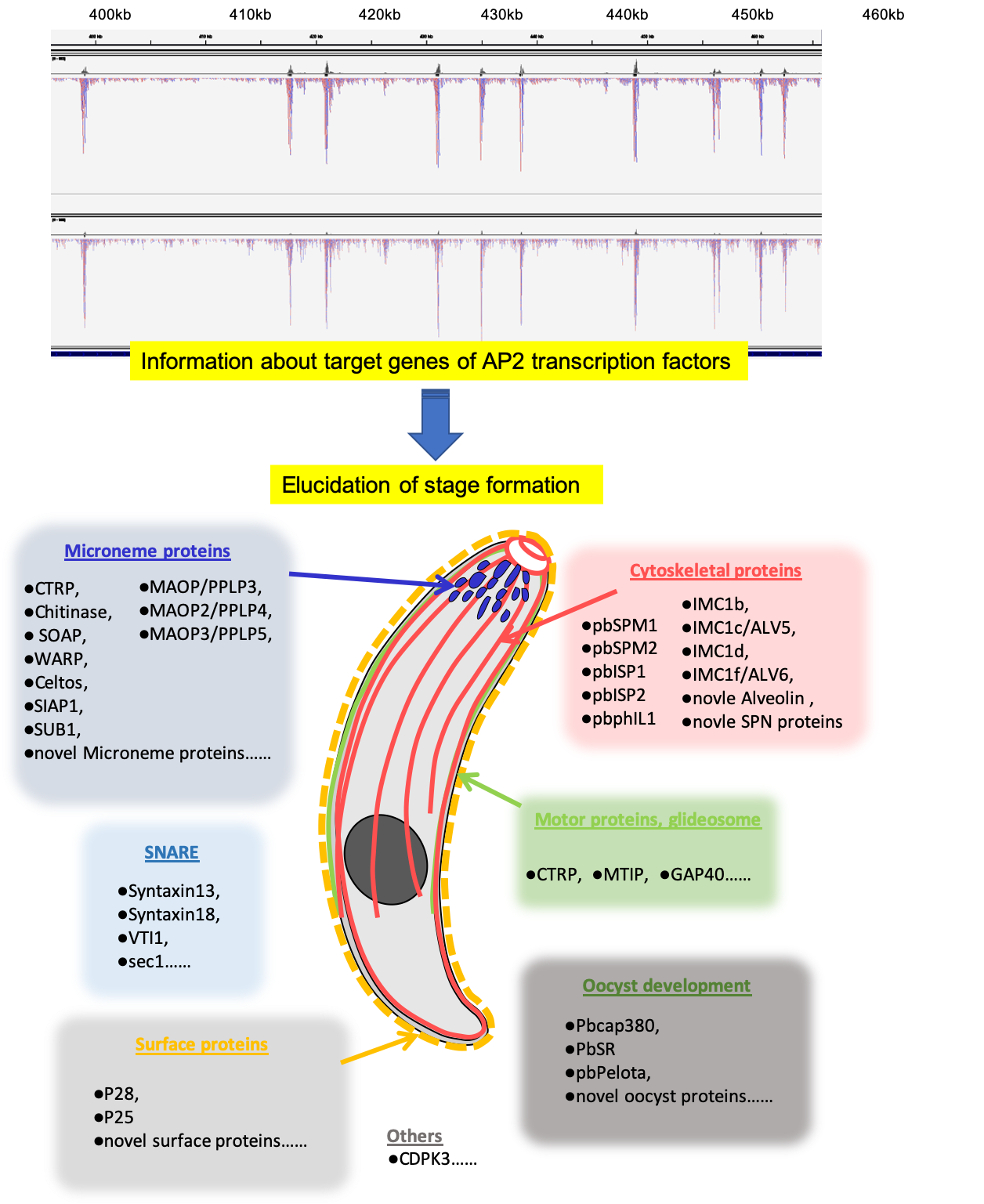
Fig. 1: Chromatin-immunoprecipitation sequencing analysis shows that AP2 transcription factor binds to the promoter region of target genes specifically. The information on target genes is useful for our understanding of the formation of the developmental stage. This figure shows the summary of the target genes in ookinete, which is the mosquito midgut invasion stage.
-
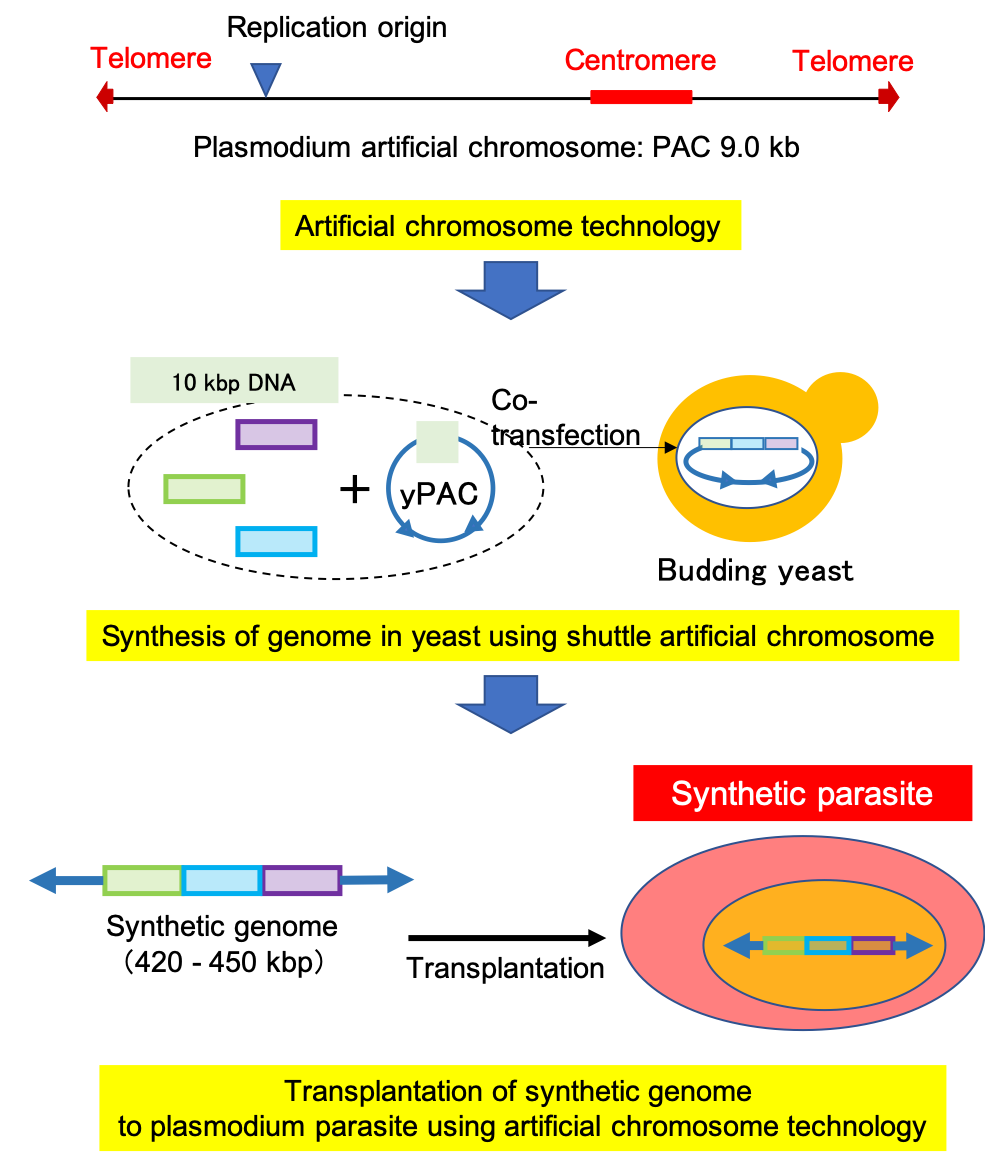
Fig. 2: Plasmodium artificial chromosome consists of a centromere, telomere, and replication origin. The shuttle artificial chromosome, yPAC, is generated by combining Plasmodium and yeast artificial chromosomes (YAC). The genome of Plasmodium parasites can be synthesized using YAC in budding yeast. Synthetic Plasmodium parasites can be generated by transplanting synthetic genome into parasites.
Staff
- Prof.: Shiroh Iwanaga
- Assoc. Prof.: Tsubasa Nishi
- Postdoc.: Mayumi Shibuya
Website
Publications
(1) Immune evasion runs in the family: two surface protein families of Plasmodium falciparum-infected erythrocytes. Chamberlain SG. et al., CurrOpin Microbiol. (2025) doi: 10.1016/j.mib.2025.102598
(2) PbARID-associated chromatin remodeling events are essential for gametocyte development in Plasmodium. Nishi T. et al., Nucleic Acids Res. (2024) 10.1093/nar/gkae207
(3) Differentiation of Plasmodium male gametocytes is initiated by the recruitment of a chromatin remodeler to a male-specific cis-element. Kaneko I. et al., Proc Natl Acad Sci U S A. (2023) 10.1073/pnas.2303432120
(4) Genome-wide functional screening of drug-resistance genes in Plasmodium falciparum. Iwanaga S. et al, Nat Commun. (2022) 10.1038/s41467-022-33804-w
- Home
- Laboratories
- Iwanaga Lab