Iwasaki Lab/International Research Center for Infectious Diseases Laboratory of Emerging Viral Diseases
Overview
The pandemic of coronavirus disease 2019 (COVID-19), caused by infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has highlighted the unprecedented significance of emerging viral infection research. In addition to SARS-CoV-2, several emerging viruses pose significant threats to the human population, but have not been well studied. Our research efforts are primarily directed toward the investigation of Lassa virus, a highly pathogenic emerging virus for which there are as yet no established therapeutic or preventive interventions. Lassa fever, a life-threatening hemorrhagic fever disease caused by Lassa virus, has garnered global attention and has been included as one of the eight Priority Diseases by the Coalition for Epidemic Preparedness Innovations (CEPI), an international organization advancing vaccine research and development. Harnessing cutting-edge techniques in virology, immunology, molecular biology, RNA biology, bioinformatics, and chemical biology, we are dedicated to understanding the mechanisms underlying virus multiplication and viral pathogenesis from various perspectives. Furthermore, we are engaged in applied research to develop novel diagnostics, treatments, and preventions based on the findings obtained from our studies.
Ongoing projects
– PolyA-independent translation regulation
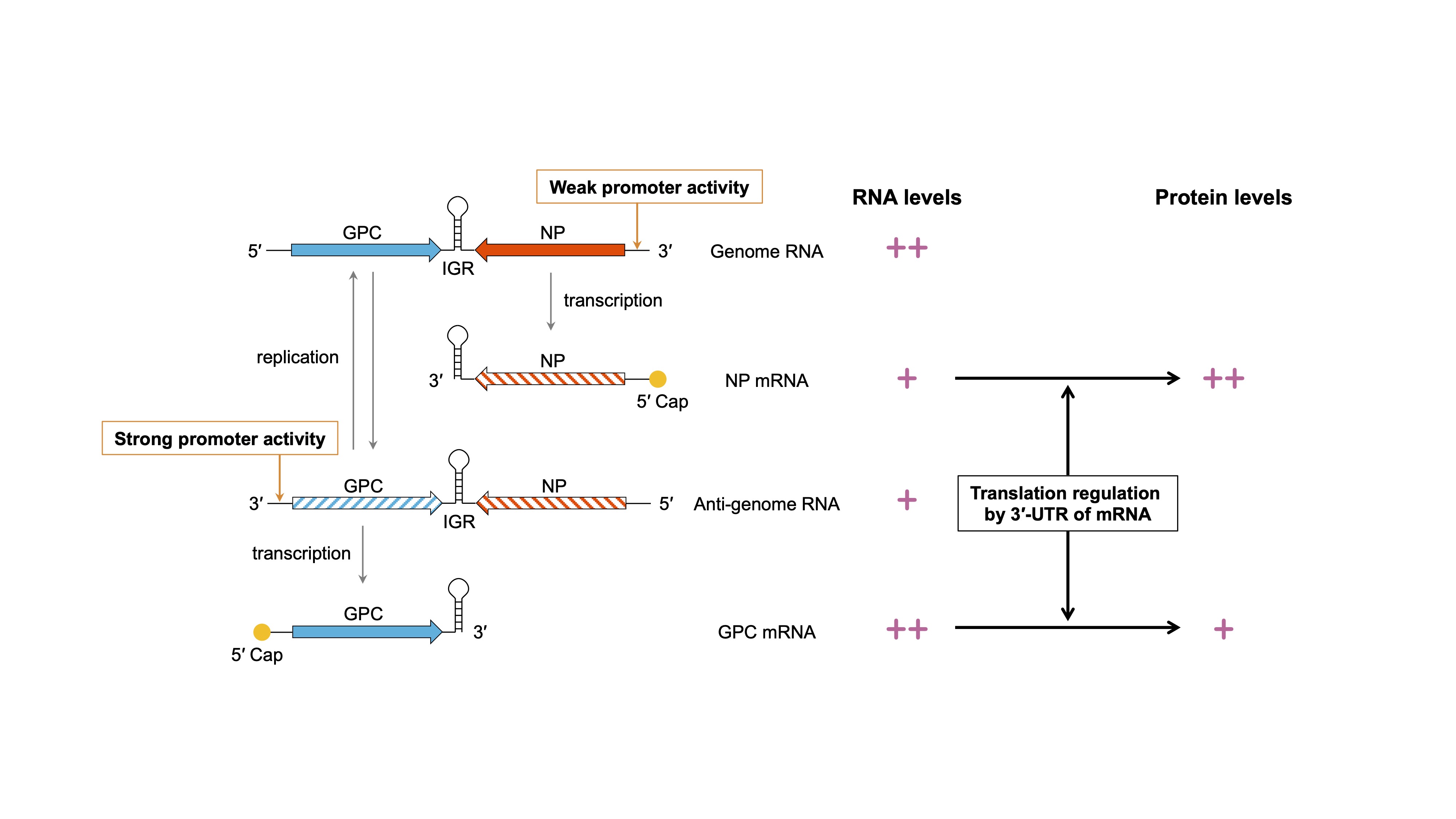
Mammalian arenaviruses (mammarenaviruses), including Lassa virus, produce mRNAs that are characterized by 5′-capped and 3′-nonpolyadenylated UTRs. We discovered that the nonpolyadenylated 3′-UTR regulates translation efficiency and identified it as a crucial mechanism underlying a characteristic coding strategy known as ambisense (Fig. 1). On the basis of this discovery, we have been working on the development of a Lassa virus live-attenuated vaccine strain that disrupts the balance of viral protein expression. Furthermore, we have identified regions (proximal regions, PRs) that are particularly important for the regulation of translation and have established a molecular foundation for the development of a novel arenavirus vector capable of controlling foreign gene expression (Fig. 2). Currently, we are involved in a more detailed investigation of the polyA-independent translation regulatory mechanism using RNA biology techniques.
– Host factors
Viruses cannot replicate on their own and require host cell machinery to achieve efficient multiplication. Leveraging genetic engineering approaches, including reverse genetics systems and the CRISPR-Cas9 system, we are engaged in the identification of host factors that contribute to Lassa virus propagation from both the viral and host perspectives. Furthermore, by understanding the host factors utilized by Lassa virus, we expect to uncover novel mechanisms involved in viral multiplication with the hope of potentially developing innovative therapeutic strategies targeting these host factors.
– Antivirals
The requirement for highly pathogenic Lassa virus to be handled in a maximum containment (biosafety level 4, BSL-4) laboratory has hampered the development of medical countermeasures against the virus. Our research has focused on conducting virus-free compound screening using a minigenome assay where Lassa virus transcription and replication activity can be assessed by reporter gene expression. Through this approach, we have identified multiple compounds that strongly inhibit Lassa virus transcription and replication. Currently, we are working towards a deeper understanding of their mechanisms of action and collaborating with experts in organic synthetic chemistry to obtain more potent Lassa virus inhibitors based on structure-activity relationships.
– Vaccines
Because cellular immunity has a major role in the recovery and prevention of disease in Lassa fever survivor cases and Lassa virus-infected animals, live-attenuated vaccines, which can confer long-term cellular and humoral immunity following a single immunization, have been considered a suitable approach for the control of Lassa virus. In contrast, in response to the COVID-19 pandemic, mRNA vaccines expressing the spike protein (SARS2-S) of SARS-CoV-2 were used worldwide. Intriguingly, studies involving humans and animals indicated that COVID-19 mRNA vaccines elicited neutralizing antibodies as well as spike protein-specific cellular immunity, suggesting that mRNA vaccines are a feasible modality against Lassa virus, where the induction of cellular immunity is crucial for virus control. Therefore, our current focus involves the investigation and advancement of novel Lassa virus mRNA vaccines.
– LCMV mouse models
Many viruses that cause diseases in humans exhibit limited infectivity in laboratory mice. However, a mammarenavirus closely related to Lassa virus, lymphocytic choriomeningitis virus (LCMV), whose natural host is Mus musculus, is one of the few human-pathogenic viruses that can efficiently infect laboratory mice without the need for genetic modifications to either the virus or mice. Thus, LCMV mouse infection models have been a powerful tool, particularly in the field of immunology, for understanding virus–host interactions, contributing to the establishment of very important immunological concepts, such as major histocompatibility complex (MHC) restriction and T cell exhaustion. Furthermore, because BSL4 facilities are not required for LCMV infection experiments, LCMV has also been used as a model virus for studying Lassa virus. We utilize various LCMV mouse infection models to unravel the mechanisms by which the virus causes disease, particularly focusing on viral infection to immune cells and innate immune responses.
– Emerging viruses
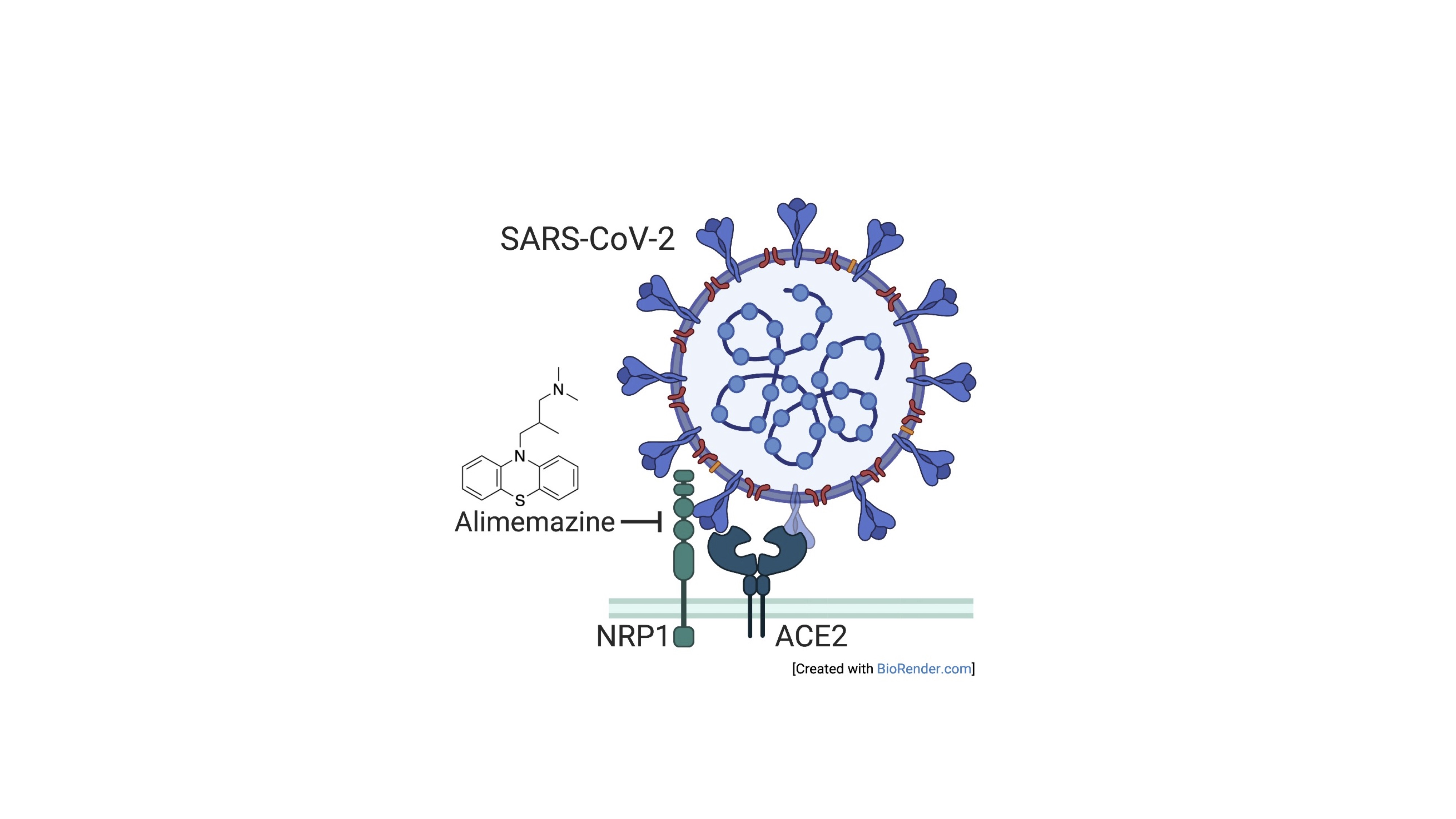
We are conducting research focused on the cell entry process of SARS-CoV-2. We demonstrated experimentally that genetic polymorphisms of the SARS-CoV-2 cell entry receptor, ACE2, initially suggested to be associated with lower fatality rates in the Asian region, have limited impact on cell entry efficiency. Additionally, through analysis using a pseudovirus that mimics the cell entry mediated by the spike protein of SARS-CoV-2, we revealed that the US Food and Drug Administration-approved drug alimemazine strongly inhibits SARS-CoV-2 infection (Fig. 3). Moreover, we intend to broaden our research scope to encompass other emerging viruses that pose potential risks to the human population, in addition to Lassa virus and SARS-CoV-2.
Staff
- SA Assoc. Prof.: Masaharu Iwasaki
- SA Asst. Prof.: Mei Hashizume
- SA Asst. Prof.: Ryotaro Nouda (concurr.)
Website
Publications
- (1) Multifunctional noncoding regions in the mammarenavirus genome. Iwasaki M., Virology. (2025) 605:110464. Review.
(2) U-73122, a phospholipase C inhibitor, impairs lymphocytic choriomeningitis virus virion infectivity. Mizuma K., Hashizume M. et al., J Gen Virol. (2024) 105:002060.
(3) An mRNA-LNP-based Lassa virus vaccine induces protective immunity in mice. Hashizume M. et al., J Virol. (2024) 98:e0057824.
(4) Phenothiazines inhibit SARS-CoV-2 cell entry via a blockade of spike protein binding to neuropilin-1. Hashizume M. et al., Antiviral Res. (2023) 209:105481.
(5) The Pan-ErbB tyrosine kinase inhibitor afatinib inhibits multiple steps of the mammarenavirus life cycle. Mizuma K. et al., Virology (2022) 576:83-95.
(6) A small stem-loop-forming region within the 3′-UTR of a non-polyadenylated LCMV mRNA promotes translation. Hashizume M. et al., J Biol Chem. (2022) 298:101576.
- Home
- Laboratories
- Iwasaki Lab