/Research Collaboration Center for Infectious Diseases Section of Viral Infections
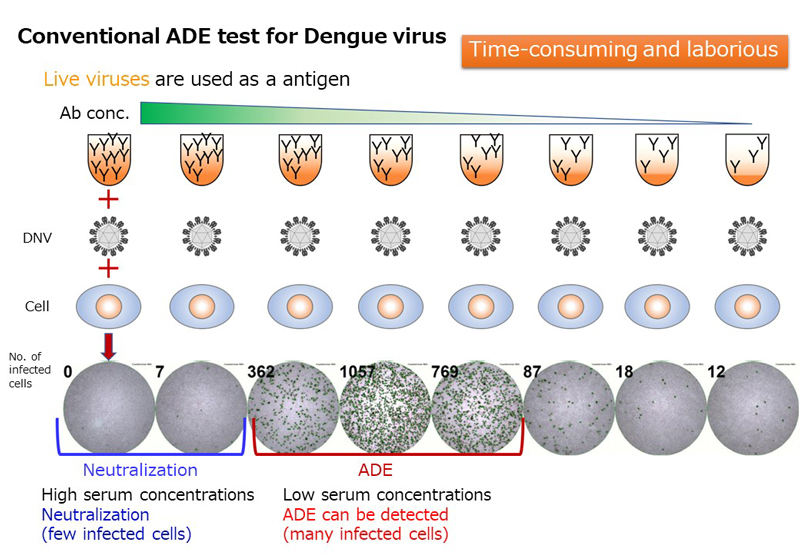
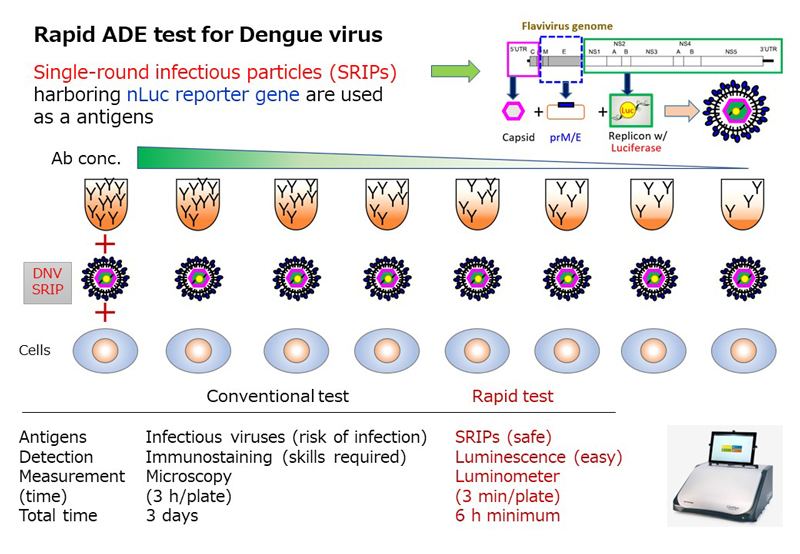
A variety of mosquito-borne viral infections are widespread in Thailand located in the tropics. Mosquito-borne viruses may spread to countries around the world, including Japan, which has strong diplomatic relations with Thailand. Therefore, establishment of countermeasures against infection of mosquito-borne viruses based on basic research is important issue in endemic prevention. Research activities focus on the antibody-dependent enhancement of infection (ADE) of dengue viruses, which are mosquito-borne viruses distributed in Thailand and are causative agent of dengue fever and severe dengue hemorrhagic fever. We are developing a rapid ADE assay, conducting cohort studies to predict the severity of dengue fever and severe dengue hemorrhagic fever, and analyzing factors associated with ADE with the aim of elucidating the mechanisms involved in the severity of these diseases.
Acute gastroenteritis caused by norovirus infection, which is observed all over the world every year, is one of the problems in public health. Viral diversification due to mutations and recombination of viral genome is thought to enable evasion from host immunity and persistent prevalence. However, it is not fully understood how viruses diversify and how they are retained in human community. Based on epidemiological surveys, we are conducting research with the aim of understanding changes in epidemic strains associated with genomic mutation and elucidating the life cycle of norovirus.
Staff
- Prof.(concur.): Takeshi Kobayashi
- SA Assoc. Prof.: Hiroto Mizushima
- SA Assoc. Prof.: Atsushi Yamanaka
Website
Publications
- (1). Antibody-dependent enhancement representing in vitro infective progeny virus titer correlates with the viremia level in dengue patients. Yamanaka A., et al., Sci Rep. (2021)11(1):12354
(2). Seroprevalence of Flavivirus Neutralizing Antibodies in Thailand by High-Throughput Neutralization Assay: Endemic Circulation of Zika Virus before 2012. Yamanaka A., et al., mSphere. (2021) 6(4):e0033921
(3). Glycan engineering of the SARS-CoV-2 receptor-binding domain elicits cross-neutralizing antibodies for SARS-related viruses. Shinnakasu R., et al., J Exp Med. (2021) 218(12):e20211003
(4). Spread of genetically similar noroviruses in Bangkok, Thailand, through symptomatic and asymptomatic individuals. Boonyos P., et al., Heliyon. (2021) 7(10):e08250
(5). Development of a Dengue Virus Serotype-Specific Non-Structural Protein 1 Capture Immunochromatography Method. Poltep K., et al., Sensors (2021) 21(23):7809
(6). The potential of COVID-19 patients' sera to cause antibody-dependent enhancement of infection and IL-6 production. Shimizu J., et al., Sci Rep.(2021) 11(1):23713
(7). Engineered flavivirus vaccines control induction of crossreactive infection-enhancing and -neutralizing antibodies. Yamanaka A., et al., Vaccine (2022) 40(42):6004-6011
(8). Japanese Encephalitis DNA Vaccines with Epitope Modification Reduce the Induction of Cross-Reactive Antibodies against Dengue Virus and Antibody-Dependent Enhancement of Dengue Virus Infection. Kotaki T., et al.,. Vaccine (2022) 10(9):1411.
(9). Development of a rapid assay system for detecting antibody-dependent enhancement of dengue virus infection. Yamanaka A., et al., J Virol Methods (2023) 311:114641
(10). Potential Protective Effect of Dengue NS1 Human Monoclonal Antibodies against Dengue and Zika Virus Infections. Sootichote R., et al., Biomedicines (2023) 11(1):227