Comparison of Culture and Culture-free Methods for Comprehensive Identification of Mycobacteria: A Single-Center Prospective Study (Nakamura Lab, in J. Clin Microbiol.)
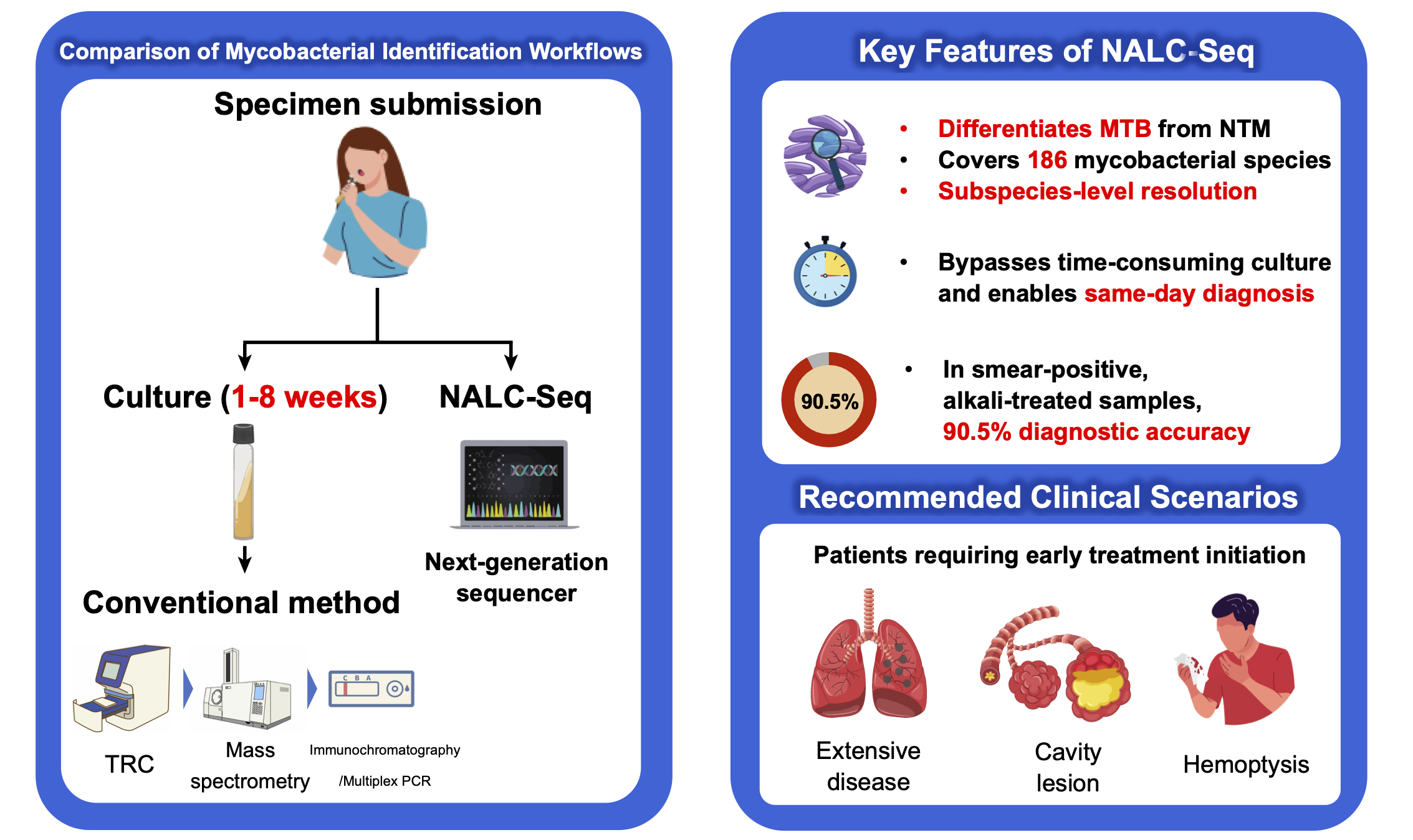
The genus Mycobacterium includes Mycobacterium tuberculosis as well as more than 200 species of nontuberculous mycobacteria (NTM). These organisms exhibit substantial diversity in clinical outcomes and drug susceptibility, making accurate species and subspecies identification essential for appropriate treatment.
However, conventional diagnostic approaches rely on culture-based methods, and the slow growth of mycobacteria often delays diagnosis by several weeks, leading to delayed treatment decisions.
In this study, we evaluated a novel culture-free approach that enables direct subspecies-level identification of mycobacteria from sputum samples using target capture sequencing.
Key Features of the Study
-
Direct analysis from sputum without the need for culture
-
Identification of over 186 mycobacterial species, including M. tuberculosis
-
Comparison with standard reference methods (culture + whole-genome sequencing)
-
Prospective cohort study conducted at Osaka Toneyama Medical Center (125 samples)
This article was published in the Journal of Clinical Microbiology, February 27, 2026
Title: Comparison of Culture and Culture-free Methods for Comprehensive Identification of Mycobacteria: A Single-Center Prospective Study”
Aurhors: Kazuki Hashimoto, Kiyoharu Fukushima, Yuki Matsumoto, Haruko Saito, Atsushi Funauchi, Nanako Hamada, June Yamauchi, Tadayoshi Nitta, Daisuke Motooka, Takuro Nii, Takanori Matsuki, Kazuyuki Tsujino, Keisuke Miki, Sho Komukai, Atsushi Kumanogoh, Shota Nakamura, Hiroshi Kida
Links
- Home
- Achievement
- Research Activities
- Comparison of Culture and Culture-free Methods for Comprehensive Identification of Mycobacteria: A Single-Center Prospective Study (Nakamura Lab, in J. Clin Microbiol.)