Kodama Lab/Division of Infectious Disease Department of Molecular Bacteriology
The increasing prevalence of antimicrobial resistance (AMR) is making infections increasingly difficult to treat, highlighting the limitations of relying solely on antibiotics for infectious disease control.
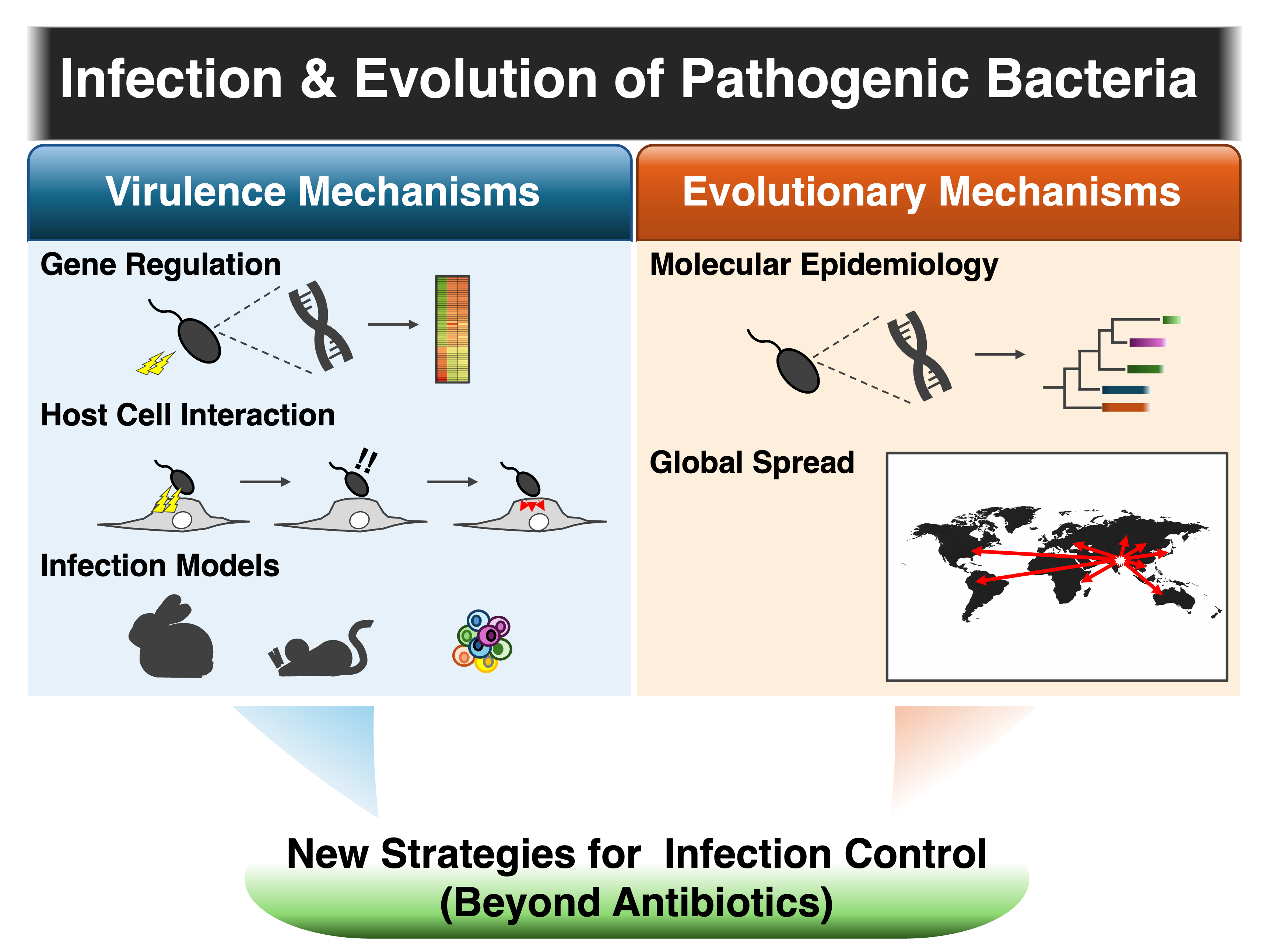
Our laboratory seeks to uncover the molecular mechanisms underlying virulence in enteric pathogenic bacteria and the evolutionary processes that drive the global emergence of these epidemic strains. By understanding these mechanisms, we aim to identify novel concepts and strategies for controlling bacterial infections beyond conventional antibiotic-based approaches.
Virulence Mechanisms in Enteropathogenic Bacteria
We investigate how enteric pathogenic bacteria express virulence and establish infection within a host. Our research integrates the identification of virulence factors with detailed analyses of their molecular functions and the regulatory mechanisms that control their expression. By combining molecular microbiology with newly developed infection models, we aim to clarify the mechanisms by which pathogens interact with the host and successfully cause disease.
Evolutionary Mechanisms Driving Global Dissemination of Epidemic Strains
Another major focus of our research is to understand why certain bacterial strains spread globally and cause large outbreaks. Through international molecular epidemiological studies, we analyze bacterial isolates collected from diverse geographic regions using whole-genome sequencing and phylogenetic approaches. By identifying lineage-specific genetic factors and experimentally validating their functions, we aim to uncover the evolutionary mechanisms that drive the emergence and global dissemination of epidemic strains.
-
Our laboratory investigates (i) molecular mechanisms of virulence in enteric pathogenic bacteria and (ii) evolutionary processes driving the global dissemination of pandemic strains. By integrating molecular microbiology, infection models, and genome-based epidemiology, we aim to develop novel strategies for infectious disease control beyond antibiotics.
Staff
- Prof.: Toshio Kodama
Publications
(1) Regulatory mechanism for host-cell contact-dependent T3SS gene expression in Vibrio parahaemolyticus. Tandhavanant S. et. al. mSystems. (2025) 10(7):e0025125.
(2) Genomic and pathogenicity analyses to identify the causative agent from multiple serogroups of non-O1, non-O139 Vibrio cholerae in foodborne outbreaks. Morita M, et. al. Microb Genom. (2025) 11(2):001364.
(3) The read-through transcription-mediated autoactivation circuit for virulence regulator expression drives robust type III secretion system 2 expression in Vibrio parahaemolyticus. Anggramukti DS, et. al. PLoS Pathog. (2024) 20(3):e1012094.
(4) Direct RNA Sequencing Unfolds the Complex Transcriptome of Vibrio parahaemolyticus. Al Kadi M, et. al. mSystems. (2021) 6(6):e0099621.
(5) Export of a Vibrio parahaemolyticus toxin by the Sec and type III secretion machineries in tandem. Matsuda S, et. al. Nature Microbiol. (2019) 4(5):781-788.
(6) VopV, an F-actin-binding type III secretion effector, is required for Vibrio parahaemolyticus-induced enterotoxicity. Hiyoshi H, et. al. Cell Host Microbe. (2011) 10(4):401-9.
- Home
- Laboratories
- Kodama Lab