Ito Lab/Bioinformatics Center Department of Biological Informatics Laboratory of Virus Informatics
The COVID-19 pandemic generated an unprecedented volume of viral genome sequences and high-throughput experimental data, ushering virology into the era of big data. Our laboratory leverages these large-scale datasets together with cutting-edge artificial intelligence (AI) technologies to develop bioinformatics and AI-based tools to control viral infectious diseases. Besides developing new computational methodologies, we aim to generate fundamental discoveries through large-scale data analyses enabled by these technologies.
AI for Predicting Viral Evolution and Its Application to Vaccine Development
The ability of viruses to evolve and alter their biological properties presents a major challenge to controlling viral infectious diseases. During the COVID-19 pandemic, successive variants emerged with enhanced immune evasion and increased infectivity, leading to greater viral fitness (i.e., transmissibility) and making epidemic control increasingly difficult.
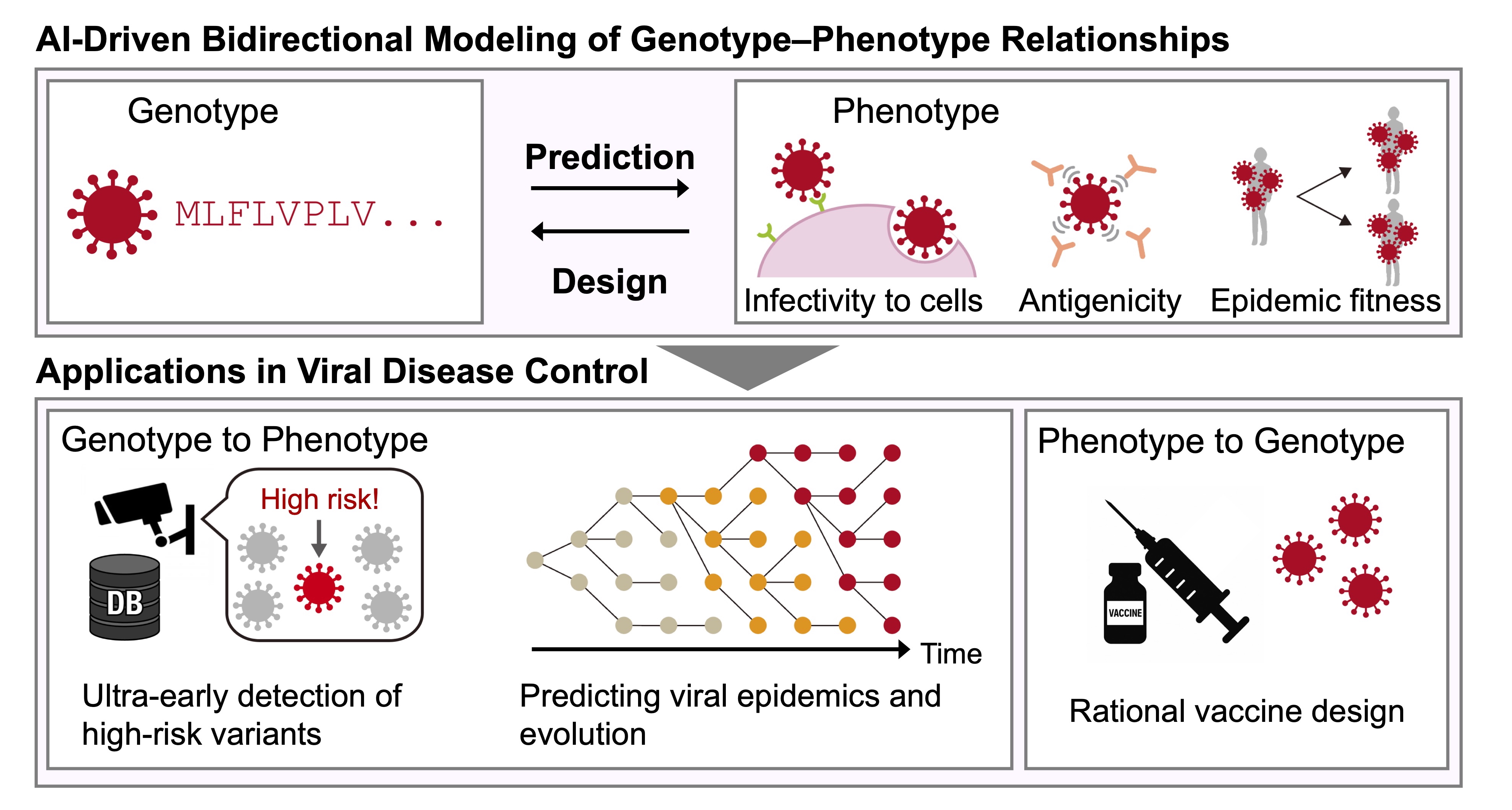
Our laboratory develops next-generation AI technologies centered on protein language models to directly predict key viral phenotypes from genome sequences, including antigenicity, cellular infectivity, and epidemic fitness. In parallel, we are constructing AI-driven design frameworks to facilitate the generation of viral proteins with specified properties, such as desired antigenic characteristics. By integrating predictive and generative approaches, we aim to enable ultra-early detection of high-risk variants, forecasting of viral epidemic dynamics and evolutionary trajectories, and rational design of vaccines and antiviral therapeutics (Fig.1).
AI for Discovering the Next Pandemic Viruses
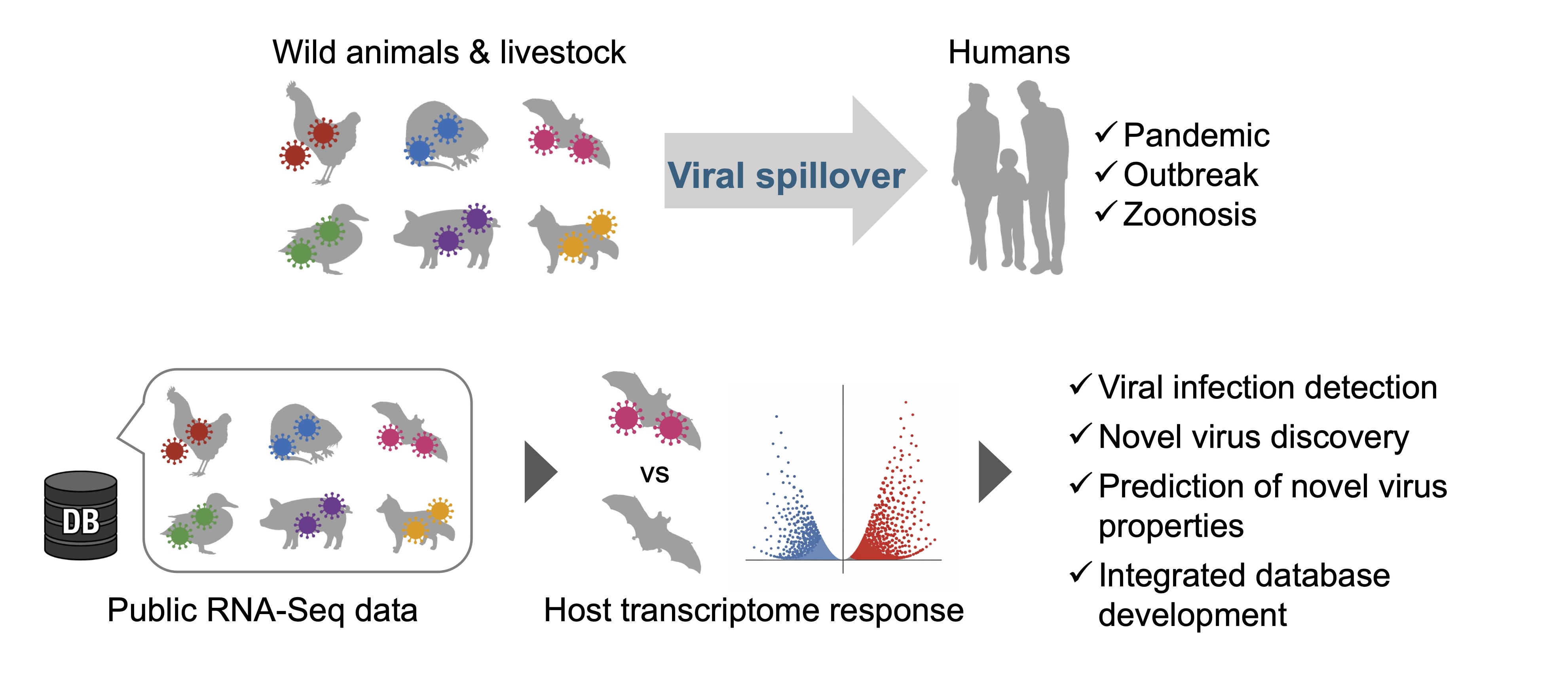
Another major factor complicating infectious disease control is zoonotic spillover, whereby virus transmission from wildlife to humans or livestock leads to the emergence of novel diseases. For example, the COVID-19 pandemic is believed to have originated from the transmission of coronaviruses circulating in horseshoe bats to humans.
Preparing for future pandemics necessitates comprehensive exploration, characterization, and cataloging of viruses circulating in wildlife and livestock populations. Our laboratory develops AI-based technologies to systematically identify novel viruses from large-scale wildlife- and livestock-derived RNA-seq datasets deposited in public databases. We further aim to infer their biological properties and estimate their pandemic risk. Specifically, we focus on developing AI models that detect viral infections based on host gene expression responses and models that predict viral phenotypes and risk potential. Through these efforts, we seek to establish a data-driven framework for pandemic preparedness grounded in large-scale genomics and AI (Fig. 2).
Staff
- Prof.: Jumpei Ito
- SA Assoc.Prof.: Junna Kawasaki
- Postdoc. : Mai Suganami
Website
Publications
(1) Lytras S et al., Inferring context-specific site variation with evotuned protein language models. NAR Genom. Bioinform. In press.
(2) Tsujino S et al., A non-spike nucleocapsid R204P mutation in SARS-CoV-2 Omicron XEC enhances inflammation and pathogenicity. Nat. Commun. (2025) 17:735.
(3) Ito J. et al., Integrative modeling of seasonal influenza evolution via AI-powered antigenic cartography. bioRxiv 2025.08.04.668423.
(4) Ito J, et al., A protein language model for exploring viral fitness landscapes. Nat. Commun. (2025) 16:4236.
(5) Tolentino J. et al., Recombination analysis on the receptor switching event of MERS-CoV and its close relatives: implications for the emergence of MERS-CoV. Virol. J. (2024) 21:84.
(6) Ito J. et al., Convergent evolution of SARS-CoV-2 Omicron subvariants leading to the emergence of BQ.1.1 variant. Nat. Commun. (2023) 14:2671.
(7) Masuda Y et al., Characterization of the evolutionary and virological aspects of mutations in the receptor binding motif of the SARS-CoV-2 spike protein. Frontiers in Virology (2023) Volume 3.
(8) Tamura T et al., Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants. Nat. Commun. (2023) 14:2800.
(9) Ito J et al., A hominoid-specific endogenous retrovirus may have rewired the gene regulatory network shared between primordial germ cells and naïve pluripotent cells. PLoS Genet. (2022) 18:e1009846.
(10) Yamasoba D et al., Virological characteristics of the SARS-CoV-2 Omicron BA.2 spike. Cell (2022) 185:2103-2115.e19.
(11) Suzuki R. et al., Attenuated fusogenicity and pathogenicity of SARS-CoV-2 Omicron variant. Nature (2022) 603:700-705.
(12) Aso H. et al., Single-cell transcriptome analysis illuminating the characteristics of species-specific innate immune responses against viral infections. Gigascience (2022) 12:giad086
(13) Kumata R. et al., A tissue level atlas of the healthy human virome. BMC. Biol. (2020) 18:55.
(14) Ito J. et al., Endogenous retroviruses drive KRAB zinc-finger protein family expression for tumor suppression. Sci. Adv. (2020) 6(43):eabc3020
(15) Ito J. et al., Retroviruses drive the rapid evolution of mammalian APOBEC3 genes. Proc. Natl. Acad. Sci. U S A (2020) 117:610-618.
- Home
- Laboratories
- Ito Lab